The lab is interested in understanding the molecular and physiological functions of mechanisms that regulate gene expression. Our current major interests are linked to the three areas described in more detail below, namely Oscillatory Gene Expression, Cell Fate Control, and MicroRNAs. However, we are happy to let the data guide us and have thus also worked at times on RNA modification, splicing, or transcription termination.
Oscillatory Gene Expression
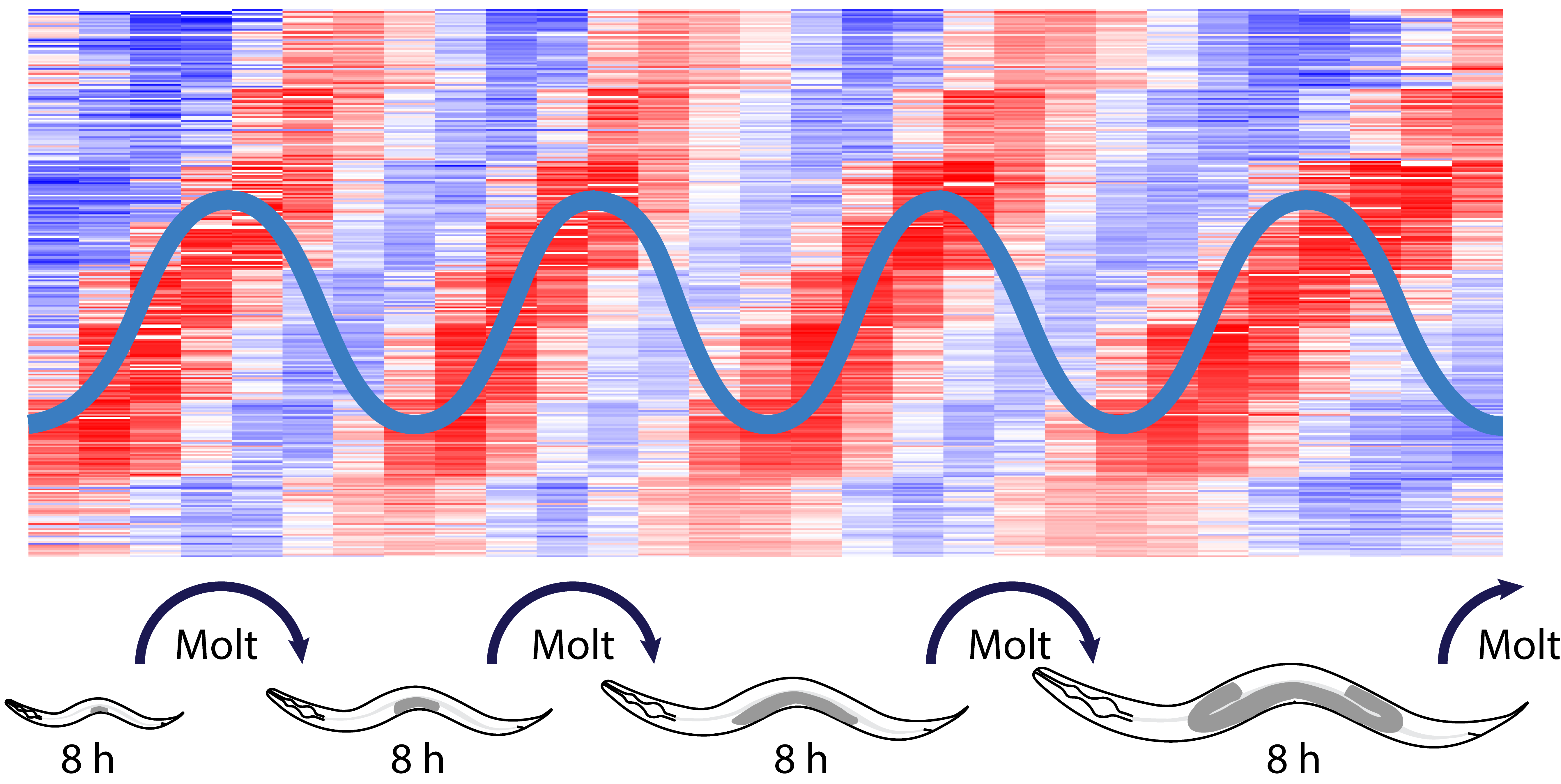
Development of a multicellular organism requires accurate regulation of events in time. Biological ‘clocks’ are thought to orchestrate such timekeeping but their molecular nature and properties have remained elusive. We discovered massive mRNA level oscillations as an apparent clock output during larval development of the nematode C. elegans (Hendriks et al., Mol Cell 2014). Characterized by a shared eight-hour period and a large amplitude (>2.1 fold), oscillations affect 20% of the active genes. These features and the accessibility of C. elegans for genetic screens make this phenomenon a powerful model to study developmental clocks. To achieve this, in a project supported by the European Resarch Council ERC, we have been investing heavily in technology to investigate oscillations and the processes they control at the single animal level. We can thus track developmental progress of animals for 384 animals at a time using a plate reader and examine development and oscillatory gene expression in parallel in tens of animals through long-term imaging. These approaches are complemented by sequencing at high temporal resolution. Using these and more standard molecular biology approaches, we have been able to show that periodic transcription is a major contributor of rhythmic mRNA accumulation, and identify candidate transcription factors. Ongoing genomic and genetic approaches aim not only at finding additional clock components and their wiring, but also at understanding their physiological role and relevance in developmental timing. Finally, by analyzing our data in the light of nonlinear dynamic theory, we have been able to obtain insight into the global architecture of the oscillator and its resulting properties (Meeuse et al., Mol Syst Biol 2020).
Cell Fate Control
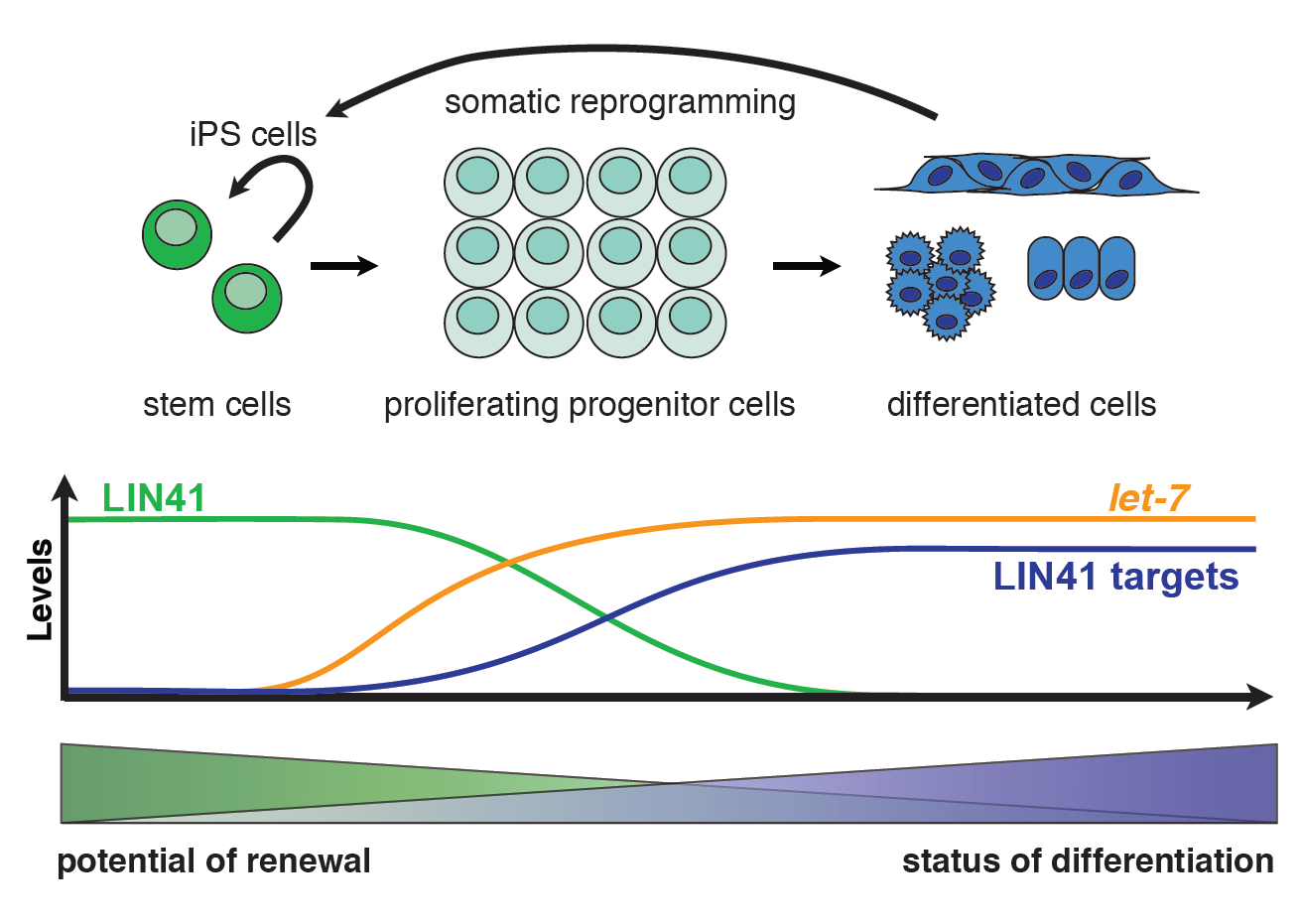
An ability to self-renew is a key feature of animal stem and progenitor cells, and therefore tightly controlled. It allows these cells to produce both a copy of themselves and a second, more differentiated daughter cell when dividing. Using C. elegans and mammalian cells, we study the mechanisms that promote self-renewal and terminate it when progenitor cells differentiate. We have a specific interest in the let-7 miRNA and its target LIN41/TRIM71, an RNA-binding protein, which regulate self-renewal and differentiation in worms as well as mammals. In C. elegans, we found that LIN41 silences its mRNA targets through distinct mechanisms, depending on the binding location: LIN41 binding to 3’UTRs promotes mRNA degradation, whereas 5’UTR-binding inhibits translation (Aeschimann et al., Mol Cell 2017). We showed that both in C. elegans and mammals, it recognizes stemloop structures with little sequence specificity (Kumari et al., Nat. Comm. 2018; Welte et al., Genes Dev 2019). We are now working on identifying the players in these distinct silencing mechanisms. We use techniques such as CRISPR-Cas9-mediated genome engineering and ribosome profiling in both C. elegans and mammalian cells with the aim of obtaining a mechanistic understanding of how LIN41 and its downstream targets control self-renewal and cell differentiation.
MicroRNAs

MicroRNAs (miRNAs) are small regulatory RNAs, ~22 nucleotides in length, that silence target mRNAs to which they bind. Through translational repression and mRNA degradation, they control diverse physiological events, and miRNA dysregulation accordingly contributes to various diseases, most notably cancers. The lab has a longstanding interest in understanding the molecular mechanisms by which miRNAs act (e.g., Ding & Großhans, EMBO J 2009), the physiological functions that they execute (e.g. Ecsedi et al., Dev Cell 2015), and the mechanisms that regulate miRNAs themselves (e.g., Chatterjee & Großhans, Nature 2009; de la Mata et al., EMBO Rep 2015). Recently, we showed that a combination of target site features and miRNA concentration determine the specificity of miRNA target recognition (Brancati & Großhans, Nucleic Acids Res 2018), and we uncovered how a protease controls small RNA loading onto Ago proteins (Gudipati et al., Mol Cell 2021).